-Search query
-Search result
Showing 1 - 50 of 60 items for (author: zhang & hq)

EMDB-35416: 
Cryo-EM structure of the Clr6S (Clr6-HDAC) complex from S. pombe
Method: single particle / : Zhang HQ, Wang X, Wang YN, Liu SM, Zhang Y, Xu K, Ji LT, Kornberg RD

EMDB-35417: 
Clr6 HDAC (Clr6S)-nucleosome complex
Method: single particle / : Zhang HQ
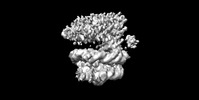
EMDB-36278: 
Rpd3S-nucleosome (H3K36me3 modified)
Method: single particle / : Wang X, Wang YN, Liu SM, Zhang Y, Xu K, Ji LT, Kornberg RD, Zhang HQ

EMDB-27205: 
Closed state of SARS-CoV-2 BA.2 variant spike protein
Method: single particle / : Zhang J, Tang WC, Gao HL, Shi W, Peng HQ, Volloch SR, Xiao TS, Chen B

EMDB-27206: 
One RBD-up state of SARS-CoV-2 BA.2 variant spike protein
Method: single particle / : Zhang J, Tang WC, Gao HL, Shi W, Peng HQ, Volloch SR, Xiao TS, Chen B

EMDB-27207: 
Middle state of SARS-CoV-2 BA.2 variant spike protein
Method: single particle / : Zhang J, Tang WC, Gao HL, Shi W, Peng HQ, Volloch SR, Xiao TS, Chen B

EMDB-29016: 
Cryo-EM structure of SARS-CoV-2 postfusion spike in membrane
Method: single particle / : Zhang J, Shi W, Cai YF, Zhu HS, Peng HQ, Voyer J, Volloch SR, Cao H, Mayer ML, Song KK, Xu C, Lu JM, Chen B
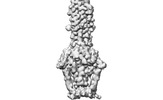
EMDB-29017: 
Cryo-EM structure of SARS-CoV-2 postfusion spike in membrane
Method: single particle / : Zhang J, Shi W, Cai YF, Zhu HS, Peng HQ, Voyer J, Volloch SR, Cao H, Mayer ML, Song KK, Xu C, Lu JM, Chen B

EMDB-29018: 
Cryo-EM structure of SARS-CoV-2 postfusion spike in membrane
Method: single particle / : Zhang J, Shi W, Cai YF, Zhu HS, Peng HQ, Voyer J, Volloch SR, Cao H, Mayer ML, Song KK, Xu C, Lu JM, Chen B

EMDB-33796: 
The NuA4 histone acetyltransferase complex from S. cerevisiae
Method: single particle / : Ji LT, Zhao LX, Xu K, Gao HH, Zhou Y, Kornberg RD, Zhang HQ

EMDB-40094: 
17b10 fab in complex with full-length SARS-CoV-2 Spike G614 trimer
Method: single particle / : Kwon HJ, Zhang J, Kosikova M, Tang WC, Rodriguez UO, Peng HQ, Meseda CA, Pedro CL, Schmeisser F, Lu JM, Zhou B, Davis CT, Wentworth DE, Chen WH, Shriver MC, Pasetti MF, Weir JP, Chen B, Xie H

EMDB-40095: 
17B10 fab in complex with up-RBD of SARS-CoV-2 Spike G614 trimer
Method: single particle / : Kwon HJ, Zhang J, Kosikova M, Tang WC, Rodriguez UO, Peng HQ, Meseda CA, Pedro CL, Schmeisser F, Lu JM, Zhou B, Davis CT, Wentworth DE, Chen WH, Shriver MC, Pasetti MF, Weir JP, Chen B, Xie H

EMDB-35522: 
Cryo-EM structure of the TUG891 bound GPR120-Giq complex(mask on receptor)
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35523: 
Cryo-EM structure of the TUG891 bound GPR120-Giq complex(mask on Giq-scFV16 complex)
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35524: 
Cryo-EM structure of the eicosapentaenoic acid bound GPR120-Gi1 complex(mask on receptor)
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35525: 
Cryo-EM structure of the eicosapentaenoic acid bound GPR120-Gi1 complex(mask on Gil-scFV16 complex)
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35529: 
Cryo-EM structure of the TUG891 bound GPR120-Giq complex (consensus map)
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35533: 
Cryo-EM structure of the eicosapentaenoic acid bound GPR120-Gi complex(consensus map)
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35356: 
Cryo-EM structure of the 9-hydroxystearic acid bound GPR120-Gi complex
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35357: 
Cryo-EM structure of the linoleic acid bound GPR120-Gi complex
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35358: 
Cryo-EM structure of the oleic acid bound GPR120-Gi complex
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35359: 
Cryo-EM structure of the TUG891 bound GPR120-Gi complex
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-35360: 
Cryo-EM structure of the eicosapentaenoic acid bound GPR120-Gi complex
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-29736: 
Cryo-EM structure of the TUG891 bound GPR120-Giq complex
Method: single particle / : Mao C, Xiao P, Tao X, Qin J, He Q, Zhang C, Yu X, Zhang Y, Sun J

EMDB-33794: 
Core module of the NuA4 complex in S. cerevisiae
Method: single particle / : Ji LT, Zhao LX, Xu K, Gao HH, Zhou Y, Kornberg RD, Zhang HQ

EMDB-32356: 
Computational design of a potent Epstein-Barr virus fusion protein gB nanoparticle vaccine
Method: subtomogram averaging / : Sun C, Wang AJ, Fang XY, Gao YZ, Liu Z, Zeng MS

EMDB-32088: 
Computational design of a potent Epstein-Barr virus fusion protein gB nanoparticle vaccine
Method: single particle / : Sun C, Fang XY, Wang AJ, Liu Z, Zeng MS
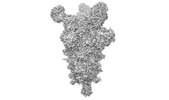
EMDB-26676: 
Three RBD-down state of SARS-CoV-2 D614G spike in complex with the SP1-77 neutralizing antibody Fab fragment
Method: single particle / : Zhang J, Luo S, Kreutzberger A, Kirchhausen T, Chen B, Haynes B, Alt F

EMDB-26677: 
Three RBD-down state of SARS-CoV-2 D614G spike in complex with the SP1-77 neutralizing antibody Fab fragment (local refinement of the RBD and Fab variable domains)
Method: single particle / : Zhang J, Luo S, Kreutzberger A, Kirchhausen T, Chen B, Haynes B, Alt F

EMDB-26678: 
An antibody from single human VH-rearranging mouse neutralizes all SARS-CoV-2 variants through BA.5 by inhibiting membrane fusion
Method: single particle / : Zhang J, Luo S, Kreutzberger A, Kirchhausen T, Chen B, Haynes B, Alt F

EMDB-27043: 
Negative stain electron microscopy single particle reconstruction of monoclonal antibody SP1-77 Fab in complex with SARS-CoV2 2P spike
Method: single particle / : Edwards RJ, Mansouri K
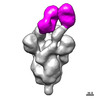
EMDB-27044: 
Negative stain electron microscopy single particle reconstruction of monoclonal antibody VHH7-7-53 Fab in complex with SARS-CoV2 2P spike
Method: single particle / : Edwards RJ, Mansouri K

EMDB-27046: 
Negative stain electron microscopy single particle reconstruction of monoclonal antibody VHH7-5-82 Fab in complex with SARS-CoV2 2P spike
Method: single particle / : Edwards RJ, Mansouri K

EMDB-26021: 
Structural and functional impact by SARS-CoV-2 Omicron spike mutations
Method: single particle / : Zhang J, Xiao TS, Cai YF, Peng HQ, Volloch SR, Chen B

EMDB-26029: 
Structural and functional impact by SARS-CoV-2 Omicron spike mutations
Method: single particle / : Zhang J, Xiao TS, Cai YF, Peng HQ, Volloch SR, Chen B

EMDB-24982: 
One RBD-up 1 of pre-fusion SARS-CoV-2 Delta variant spike protein
Method: single particle / : Zhang J, Xiao TS, Cai YF, Peng HQ, Volloch SR, Chen B

EMDB-24988: 
One RBD-up 2 of pre-fusion SARS-CoV-2 Gamma variant spike protein
Method: single particle / : Zhang J, Xiao TS, Cai YF, Peng HQ, Volloch SR, Chen B

EMDB-24981: 
Closed state of pre-fusion SARS-CoV-2 Delta variant spike protein
Method: single particle / : Zhang J, Xiao TS, Cai YF, Peng HQ, Volloch SR, Chen B

EMDB-24983: 
One RBD-up 2 of pre-fusion SARS-CoV-2 Delta variant spike protein
Method: single particle / : Zhang J, Xiao TS, Cai YF, Peng HQ, Volloch SR, Chen B

EMDB-24984: 
Closed state of pre-fusion SARS-CoV-2 Kappa variant spike protein
Method: single particle / : Zhang J, Xiao TS, Cai YF, Peng HQ, Volloch SR, Chen B

EMDB-24985: 
One RBD-up 1 of pre-fusion SARS-CoV-2 Kappa variant spike protein
Method: single particle / : Zhang J, Xiao TS, Cai YF, Peng HQ, Volloch SR, Chen B

EMDB-24986: 
One RBD-up 2 of pre-fusion SARS-CoV-2 Kappa variant spike protein
Method: single particle / : Zhang J, Xiao TS, Cai YF, Peng HQ, Volloch SR, Chen B

EMDB-24987: 
One RBD-up 1 of pre-fusion SARS-CoV-2 Gamma variant spike protein
Method: single particle / : Zhang J, Xiao TS, Cai YF, Peng HQ, Volloch SR, Chen B

EMDB-24121: 
Structural basis for enhanced infectivity and immune evasion of SARS-CoV-2 variants
Method: single particle / : Zhang J, Cai YF, Xiao TS, Rawson S, Peng HQ, Sterling SM, Walsh Jr RM, Volloch SR, Chen B

EMDB-24122: 
Structural basis for enhanced infectivity and immune evasion of SARS-CoV-2 variants
Method: single particle / : Zhang J, Cai YF, Xiao TS, Rawson S, Peng HQ, Sterling SM, Walsh Jr RM, Volloch SR, Chen B

EMDB-24123: 
Structural basis for enhanced infectivity and immune evasion of SARS-CoV-2 variants
Method: single particle / : Zhang J, Cai YF, Xiao TS, Rawson S, Peng HQ, Sterling SM, Walsh Jr RM, Volloch SR, Chen B

EMDB-24124: 
Structural basis for enhanced infectivity and immune evasion of SARS-CoV-2 variants
Method: single particle / : Zhang J, Cai YF, Xiao TS, Rawson S, Peng HQ, Sterling SM, Walsh Jr RM, Volloch SR, Chen B

EMDB-24125: 
Structural basis for enhanced infectivity and immune evasion of SARS-CoV-2 variants
Method: single particle / : Zhang J, Cai YF, Xiao TS, Rawson S, Peng HQ, Sterling SM, Walsh Jr RM, Volloch SR, Chen B

EMDB-24126: 
Structural basis for enhanced infectivity and immune evasion of SARS-CoV-2 variants
Method: single particle / : Zhang J, Cai YF, Xiao TS, Rawson S, Peng HQ, Sterling SM, Walsh Jr RM, Volloch SR, Chen B

EMDB-24127: 
Structural basis for enhanced infectivity and immune evasion of SARS-CoV-2 variants
Method: single particle / : Zhang J, Cai YF, Xiao TS, Rawson S, Peng HQ, Sterling SM, Walsh Jr RM, Volloch SR, Chen B
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
